遗传算法求TSP问题
目录
人工智能第四次实验报告 1
遗传算法求TSP问题 1
一 、问题背景 1
1.1 遗传算法简介 1
1.2 遗传算法基本要素 2
1.3 遗传算法一般步骤 2
二 、程序说明 3
2.3 选择初始群体 4
2.4 适应度函数 4
2.5 遗传操作 4
2.6 迭代过程 4
三 、程序测试 5
3.1 求解不同规模的TSP问题的算法性能 5
3.2 种群规模对算法结果的影响 5
3.3 交叉概率对算法结果的影响 6
3.4 变异概率对算法结果的影响 7
3.5 交叉概率和变异概率对算法结果的影响 7
四 、算法改进 8
4.1 块逆转变异策略 8
4.2 锦标赛选择法 9
五 、实验总结 10
一 、问题背景
1.1遗传算法简介
遗传算法是一种进化算法,基于自然选择和生物遗传等生物进化机制的一种搜索算法,其通过选 择、重组和变异三种操作实现优化问题的求解。它的本质是从原问题的一组解出发改进到另一组较好的 解,再从这组改进的解出发进一步改进。在搜索过程中,它利用结构和随机的信息,是满足目标的决策 获得最大的生存可能,是一种概率型算法。
遗传算法主要借用生物中“适者生存”的原则,在遗传算法中,染色体对应的是数据或数组,通常由 一维的串结构数据来表示。串上的各个位置对应一个基因座,而各个位置上所取的值对等位基因。遗传 算法处理的是基因型个体,一定数量的个体组成了群体。群体的规模就是个体的数目。不同个体对环境 的适应度不同,适应度打的个体被选择进行遗传操作产生新个体。本文转载自http://www.biyezuopin.vip/onews.asp?id=16719每次选择两个染色体进行产生一组新 染色体,染色体也可能发生变异,得到下一代群体。
1.2遗传算法基本要素
1.参数编码:可以采用位串编码、实数编码、多参数级联编码等
2.设定初始群体:
1.启发 / 非启发给定一组解作为初始群体
2.确定初始群体的规模
3.设定适应度函数:将目标函数映射为适应度函数,可以进行尺度变换来保证非负、归一等特性
4.设定遗传操作:
1.选择:从当前群体选出一系列优良个体,让他们产生后代个体
2.交叉:两个个体的基因进行交叉重组来获得新个体
3.变异:随机变动个体串基因座上的某些基因
5.设定控制参数:例如变异概率、交叉程度、迭代上限等。
import numpy as np
import random
import matplotlib.pyplot as plt
import copy
import timefrom matplotlib.ticker import MultipleLocator
from scipy.interpolate import interpolateCITY_NUM = 20
City_Map = 100 * np.random.rand(CITY_NUM, 2)DNA_SIZE = CITY_NUM #编码长度
POP_SIZE = 100 #种群大小
CROSS_RATE = 0.6 #交叉率
MUTA_RATE = 0.2 #变异率
Iterations = 1000 #迭代次数# 根据DNA的路线计算距离
def distance(DNA):dis = 0temp = City_Map[DNA[0]]for i in DNA[1:]:dis = dis + ((City_Map[i][0]-temp[0])**2+(City_Map[i][1]-temp[1])**2)**0.5temp = City_Map[i]return dis+((temp[0]-City_Map[DNA[0]][0])**2+(temp[1]-City_Map[DNA[0]][1])**2)**0.5# 计算种群适应度,这里适应度用距离的倒数表示
def getfitness(pop):temp = []for i in range(len(pop)):temp.append(1/(distance(pop[i])))return temp-np.min(temp) + 0.000001# 选择:根据适应度选择,以赌轮盘的形式,适应度越大的个体被选中的概率越大
def select(pop, fitness):s = fitness.sum()temp = np.random.choice(np.arange(len(pop)), size=POP_SIZE, replace=True,p=(fitness/s))p = []for i in temp:p.append(pop[i])return p# 4.2 选择:锦标赛选择法
def selectII(pop, fitness):p = []for i in range(POP_SIZE):temp1 = np.random.randint(POP_SIZE)temp2 = np.random.randint(POP_SIZE)DNA1 = pop[temp1]DNA2 = pop[temp2]if fitness[temp1] > fitness[temp2]:p.append(DNA1)else:p.append(DNA2)return p# 变异:选择两个位置互换其中的城市编号
def mutation(DNA, MUTA_RATE):if np.random.rand() < MUTA_RATE: # 以MUTA_RATE的概率进行变异# 随机产生两个实数,代表要变异基因的位置,确保两个位置不同,将2个所选位置进行互换mutate_point1 = np.random.randint(0, DNA_SIZE)mutate_point2 = np.random.randint(0,DNA_SIZE)while(mutate_point1 == mutate_point2):mutate_point2 = np.random.randint(0,DNA_SIZE)DNA[mutate_point1],DNA[mutate_point2] = DNA[mutate_point2],DNA[mutate_point1]# 4.1 变异:在父代中随机选择两个点,然后反转之间的部分
def mutationII(DNA, MUTA_RATE):if np.random.rand() < MUTA_RATE:mutate_point1 = np.random.randint(0, DNA_SIZE)mutate_point2 = np.random.randint(0, DNA_SIZE)while (mutate_point1 == mutate_point2):mutate_point2 = np.random.randint(0, DNA_SIZE)if(mutate_point1 > mutate_point2):mutate_point1, mutate_point2 = mutate_point2, mutate_point1DNA[mutate_point1:mutate_point2].reverse()# 4.1 变异:调用 I 和 II
def mutationIII(DNA, MUTA_RATE):mutationII(DNA, MUTA_RATE)mutation(DNA, MUTA_RATE)# 交叉变异
# muta = 1时变异调用 mutation;
# muta = 2时变异调用 mutationII;
# muta = 3时变异调用 mutationIII
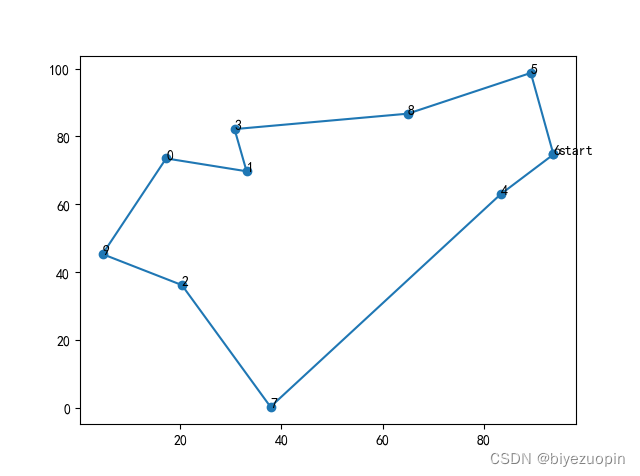
def crossmuta(pop, CROSS_RATE, muta=1):new_pop = []for i in range(len(pop)): # 遍历种群中的每一个个体,将该个体作为父代n = np.random.rand()if n >= CROSS_RATE: # 大于交叉概率时不发生变异,该子代直接进入下一代temp = pop[i].copy()new_pop.append(temp)# 小于交叉概率时发生变异if n < CROSS_RATE:# 选取种群中另一个个体进行交叉list1 = pop[i].copy()list2 = pop[np.random.randint(POP_SIZE)].copy()status = True# 产生2个不相等的节点,中间部分作为交叉段,采用部分匹配交叉while status:k1 = random.randint(0, len(list1) - 1)k2 = random.randint(0, len(list2) - 1)if k1 < k2:status = Falsek11 = k1# 两个DNA中待交叉的片段fragment1 = list1[k1: k2]fragment2 = list2[k1: k2]# 交换片段后的DNAlist1[k1: k2] = fragment2list2[k1: k2] = fragment1# left1就是 list1除去交叉片段后剩下的DNA片段del list1[k1: k2]left1 = list1offspring1 = []for pos in left1:# 如果 left1 中有与待插入的新片段相同的城市编号if pos in fragment2:# 找出这个相同的城市编号在在原DNA同位置编号的位置的城市编号# 循环查找,直至这个城市编号不再待插入的片段中pos = fragment1[fragment2.index(pos)]while pos in fragment2:pos = fragment1[fragment2.index(pos)]# 修改原DNA片段中该位置的城市编号为这个新城市编号offspring1.append(pos)continueoffspring1.append(pos)for i in range(0, len(fragment2)):offspring1.insert(k11, fragment2[i])k11 += 1temp = offspring1.copy()# 根据 type 的值选择一种变异策略if muta == 1:mutation(temp, MUTA_RATE)elif muta == 2:mutationII(temp, MUTA_RATE)elif muta == 3:mutationIII(temp, MUTA_RATE)# 把部分匹配交叉后形成的合法个体加入到下一代种群new_pop.append(temp)return new_popdef print_info(pop):fitness = getfitness(pop)maxfitness = np.argmax(fitness) # 得到种群中最大适应度个体的索引print("最优的基因型:", pop[maxfitness])print("最短距离:",distance(pop[maxfitness]))# 按最优结果顺序把地图上的点加入到best_map列表中best_map = []for i in pop[maxfitness]:best_map.append(City_Map[i])best_map.append(City_Map[pop[maxfitness][0]])X = np.array((best_map))[:,0]Y = np.array((best_map))[:,1]# 绘制地图以及路线plt.figure()plt.rcParams['font.sans-serif'] = ['SimHei']plt.scatter(X,Y)for dot in range(len(X)-1):plt.annotate(pop[maxfitness][dot],xy=(X[dot],Y[dot]),xytext = (X[dot],Y[dot]))plt.annotate('start',xy=(X[0],Y[0]),xytext = (X[0]+1,Y[0]))plt.plot(X,Y)# 3.2 种群规模对算法结果的影响
def pop_size_test():global POP_SIZEITE = 3 # 每个值测试多次求平均数以降低随机误差i_list = [10, 50, 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000]b_list = []t_list = []for i in i_list:print(i)POP_SIZE = itime_cost = 0min_path = 0for j in range(ITE):time_start = time.time()ans = tsp_solve()min_path += min(ans)time_end = time.time()time_cost += time_end - time_startb_list.append(min_path / ITE)t_list.append(time_cost / ITE)show_test_result(i_list, b_list, t_list, "POP_SIZE")# 3.3 交叉概率对算法结果的影响
def cross_rate_test():global CROSS_RATEITE = 3 # 每个值测试多次求平均数以降低随机误差i_list = range(0, 21)b_list = []t_list = []ii_list = [] # [0, 0.05, 0.1, ... 0.95, 1]for i in i_list:print(i)CROSS_RATE = 0.05 * iii_list.append(CROSS_RATE)time_cost = 0min_path = 0for j in range(ITE):time_start = time.time()ans = tsp_solve()min_path += min(ans)time_end = time.time()time_cost += time_end - time_startb_list.append(min_path / ITE)t_list.append(time_cost / ITE)show_test_result(ii_list, b_list, t_list, "CROSS_RATE")# 3.4 变异概率对算法结果的影响
def muta_rate_test():global MUTA_RATEITE = 3 # 每个值测试多次求平均数以降低随机误差i_list = range(0, 21)b_list = []t_list = []ii_list = [] # [0, 0.05, 0.1, ... 0.95, 1]for i in i_list:print(i)MUTA_RATE = 0.05 * iii_list.append(MUTA_RATE)time_cost = 0min_path = 0for j in range(ITE):time_start = time.time()ans = tsp_solve()min_path += min(ans)time_end = time.time()time_cost += time_end - time_startb_list.append(min_path / ITE)t_list.append(time_cost / ITE)show_test_result(ii_list, b_list, t_list, "MUTA_RATE")# 3.5 交叉概率和变异概率对算法结果的影响
def cross_muta_test():s = np.array([0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0])X, Y = np.meshgrid(s,s)Z = np.zeros(shape=(11, 11))global MUTA_RATEglobal CROSS_RATEfor i in range(11):for j in range(11):print(str(i) + ":" + str(j))CROSS_RATE = X[0,i]MUTA_RATE = Y[0,j]ans = tsp_solve()Z[i, j] = min(ans)ax = plt.axes(projection='3d')ax.plot_surface(X, Y, Z, rstride=1, cstride=1,cmap='rainbow', edgecolor='none')ax.set_xlabel("CROSS_RATE")ax.set_ylabel("MUTA_RATE")ax.set_zlabel("Shortest_Path")ax.set_title('TSP')plt.show()# 3.2-3.4 生成参数测试结果的可视化图表
def show_test_result(i_list, b_list, t_list, msg):ax1 = plt.subplot(121)ax1.plot(i_list, b_list, 'b')ax1.set_xlabel(msg)ax1.set_ylabel("Shortest Path")ax2 = plt.subplot(122)ax2.plot(i_list, t_list, 'r')ax2.set_xlabel(msg)ax2.set_ylabel("Cost Time")plt.show()# 求解TSP问题并返回最大值
# muta 指定变异方式,sel 指定选择方式
def tsp_solve(muta=1, sel=1):pop = []li = list(range(DNA_SIZE))for i in range(POP_SIZE):random.shuffle(li)l = li.copy()pop.append(l)best_dis = []# 进行选择,交叉,变异,并把每代的最优个体保存在best_dis中for i in range(Iterations): # 迭代N代pop = crossmuta(pop, CROSS_RATE, muta=muta)fitness = getfitness(pop)maxfitness = np.argmax(fitness)best_dis.append(distance(pop[maxfitness]))if sel == 1:pop = select(pop, fitness) # 选择生成新的种群elif sel == 2:pop = selectII(pop, fitness) # 选择生成新的种群return best_dis# 4.1 块逆转变异策略对比测试
def opt1_test():ITE = 20 # 测试次数i_list = range(ITE)b_list = [] # 每次求出的最短路径t_list = [] # 每次求解的耗时b_listII = []t_listII = []b_listIII = []t_listIII = []for i in i_list:print(i)# I. 原两点互换异策略time_start = time.time()b_list.append(min(tsp_solve(muta=1)))time_end = time.time()t_list.append(time_end - time_start)# II. 块逆转变异策略time_startII = time.time()b_listII.append(min(tsp_solve(muta=2)))time_endII = time.time()t_listII.append(time_endII - time_startII)# III. 同时使用上述两种编译策略time_startIII = time.time()b_listIII.append(min(tsp_solve(muta=3)))time_endIII = time.time()t_listIII.append(time_endIII - time_startIII)# 做排序处理,方便比较b_list.sort()t_list.sort()b_listII.sort()t_listII.sort()b_listIII.sort()t_listIII.sort()ax1 = plt.subplot(121)ax1.plot(i_list, b_list, 'b', label="Origin")ax1.plot(i_list, b_listII, 'r', label="Block-reversal")ax1.plot(i_list, b_listIII, 'g', label="Origin + Block-reversal")ax1.set_ylabel("Shortest Path")ax2 = plt.subplot(122)ax2.plot(i_list, t_list, 'b', label="Origin")ax2.plot(i_list, t_listII, 'r', label="Block-reversal")ax2.plot(i_list, t_listIII, 'g', label="Origin + Block-reversal")ax2.set_ylabel("Cost Time")plt.legend()plt.show()# 4.2 锦标赛选择策略对比测试
def opt2_test():ITE = 20 # 测试次数i_list = range(ITE)b_list = [] # 每次求出的最短路径t_list = [] # 每次求解的耗时b_listII = []t_listII = []b_listIII = []t_listIII = []for i in i_list:print(i)# I. 原赌轮盘选择策略time_start = time.time()b_list.append(min(tsp_solve(sel=1)))time_end = time.time()t_list.append(time_end - time_start)# II. 锦标赛选择策略time_startII = time.time()b_listII.append(min(tsp_solve(sel=2)))time_endII = time.time()t_listII.append(time_endII - time_startII)# III. 锦标赛选择策略 + 两点互换变异 + 块逆转变异策略time_startIII = time.time()b_listIII.append(min(tsp_solve(sel=2,muta=3)))time_endIII = time.time()t_listIII.append(time_endIII - time_startIII)# 做排序处理,方便比较b_list.sort()t_list.sort()b_listII.sort()t_listII.sort()b_listIII.sort()t_listIII.sort()ax1 = plt.subplot(121)ax1.plot(i_list, b_list, 'b', label="Origin")ax1.plot(i_list, b_listII, 'r', label="Tournament")ax1.plot(i_list, b_listIII, 'g', label="Tournament + Block-reversal + Origin")ax1.set_ylabel("Shortest Path")ax2 = plt.subplot(122)ax2.plot(i_list, t_list, 'b', label="Origin")ax2.plot(i_list, t_listII, 'r', label="Tournament")ax2.plot(i_list, t_listIII, 'g', label="Tournament + Block-reversal + Origin")ax2.set_ylabel("Cost Time")plt.legend()plt.show()# 3.1 原程序的主函数 - 求解不同规模的TSP问题的算法性能
def ori_main():time_start = time.time()pop = [] # 生成初代种群popli = list(range(DNA_SIZE))for i in range(POP_SIZE):random.shuffle(li)l = li.copy()pop.append(l)best_dis= []# 进行选择,交叉,变异,并把每代的最优个体保存在best_dis中for i in range(Iterations): # 迭代N代pop = crossmuta(pop, CROSS_RATE)fitness = getfitness(pop)maxfitness = np.argmax(fitness)best_dis.append(distance(pop[maxfitness]))pop = select(pop, fitness) # 选择生成新的种群time_end = time.time()print_info(pop)print('逐代的最小距离:',best_dis)print('Totally cost is', time_end - time_start, "s")plt.figure()plt.plot(range(Iterations),best_dis)# 4.1 块逆转变异策略运行效果展示
def opt1_main():time_start = time.time()pop = [] # 生成初代种群popli = list(range(DNA_SIZE))for i in range(POP_SIZE):random.shuffle(li)l = li.copy()pop.append(l)best_dis= []# 进行选择,交叉,变异,并把每代的最优个体保存在best_dis中for i in range(Iterations): # 迭代N代pop = crossmuta(pop, CROSS_RATE, muta=3)fitness = getfitness(pop)maxfitness = np.argmax(fitness)best_dis.append(distance(pop[maxfitness]))pop = select(pop, fitness) # 选择生成新的种群time_end = time.time()print_info(pop)print('逐代的最小距离:',best_dis)print('Totally cost is', time_end - time_start, "s")plt.figure()plt.plot(range(Iterations),best_dis)if __name__ == "__main__":ori_main() # 原程序的主函数opt1_main() # 块逆转变异策略运行效果展示plt.show()plt.close()# opt1_test() # 块逆转变异策略对比测试# opt2_test() # 锦标赛选择策略对比测试# pop_size_test() # POP_SIZE 种群规模参数测试# cross_rate_test() # CROSS_RATE 交叉率参数测试# muta_rate_test() # MUTA_RATE 变异率参数测试# cross_muta_test() # 交叉率和变异率双参数测试