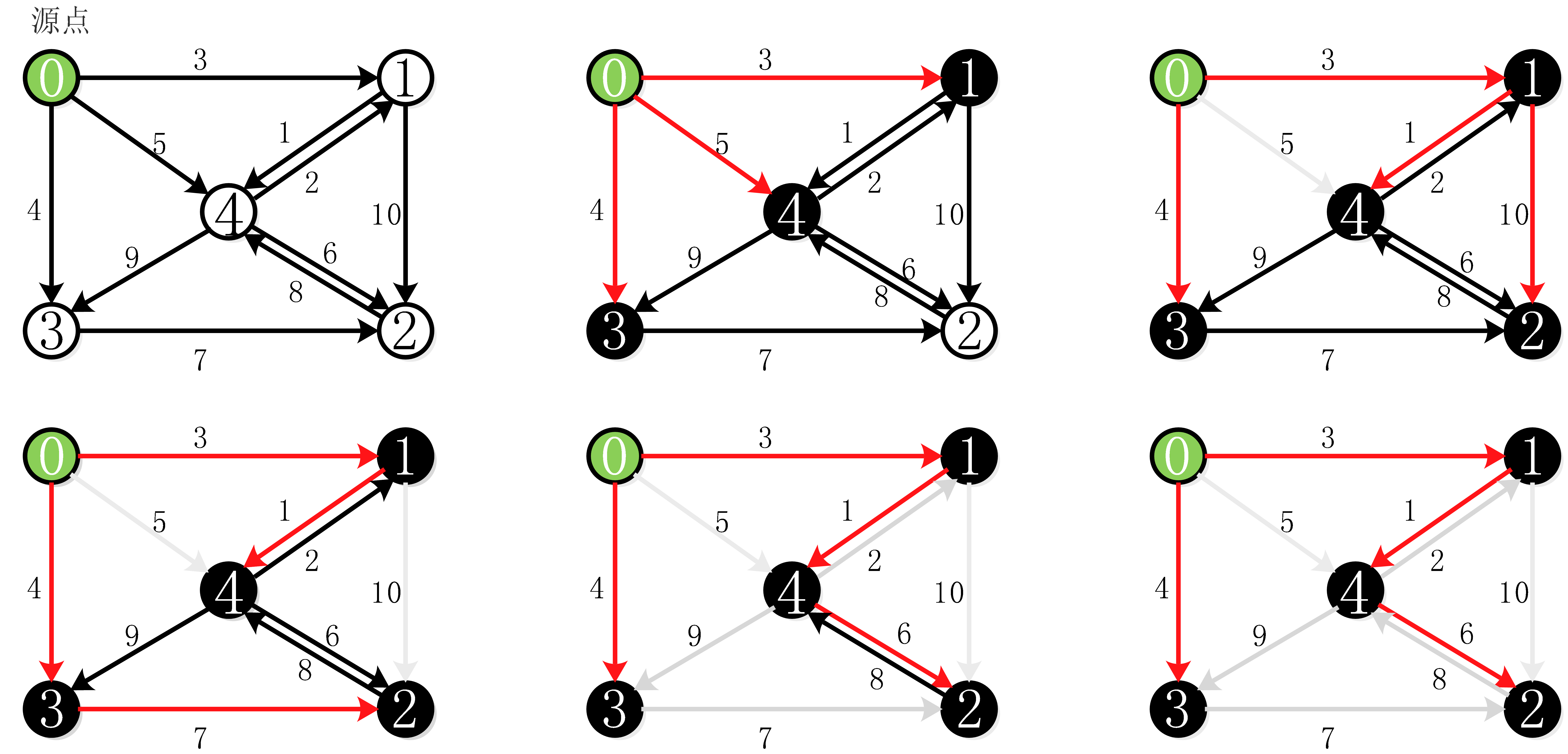
NSGA-Ⅲ源代码如下,供大家学习和应用。该算法在梯级水电-火电的应用订阅专栏即可查看:

1、主函数
%
% Copyright (c) 2016, Mostapha Kalami Heris & Yarpiz (www.yarpiz.com)
% All rights reserved. Please read the "LICENSE" file for license terms.
%
% Project Code: YPEA126
% Project Title: Non-dominated Sorting Genetic Algorithm III (NSGA-III)
% Publisher: Yarpiz (www.yarpiz.com)
%
% Implemented by: Mostapha Kalami Heris, PhD (member of Yarpiz Team)
%
% Cite as:
% Mostapha Kalami Heris, NSGA-III: Non-dominated Sorting Genetic Algorithm, the Third Version — MATLAB Implementation (URL: https://yarpiz.com/456/ypea126-nsga3), Yarpiz, 2016.
%
% Contact Info: sm.kalami@gmail.com, info@yarpiz.com
%
% Base Reference Paper:
% K. Deb and H. Jain, "An Evolutionary Many-Objective Optimization Algorithm
% Using Reference-Point-Based Nondominated Sorting Approach, Part I: Solving
% Problems With Box Constraints, "
% in IEEE Transactions on Evolutionary Computation,
% vol. 18, no. 4, pp. 577-601, Aug. 2014.
%
% Reference Paper URL: http://doi.org/10.1109/TEVC.2013.2281535
% nsga3;
2、NSGA-Ⅲ函数
clc;
clear;
close all;%% Problem DefinitionCostFunction = @(x) MOP2(x); % Cost FunctionnVar = 5; % Number of Decision VariablesVarSize = [1 nVar]; % Size of Decision Variables MatrixVarMin = -1; % Lower Bound of Variables
VarMax = 1; % Upper Bound of Variables% Number of Objective Functions
nObj = numel(CostFunction(unifrnd(VarMin, VarMax, VarSize)));%% NSGA-II Parameters% Generating Reference Points
nDivision = 10;
Zr = GenerateReferencePoints(nObj, nDivision);MaxIt = 50; % Maximum Number of IterationsnPop = 80; % Population SizepCrossover = 0.5; % Crossover Percentage
nCrossover = 2*round(pCrossover*nPop/2); % Number of Parnets (Offsprings)pMutation = 0.5; % Mutation Percentage
nMutation = round(pMutation*nPop); % Number of Mutantsmu = 0.02; % Mutation Ratesigma = 0.1*(VarMax-VarMin); % Mutation Step Size%% Colect Parametersparams.nPop = nPop;
params.Zr = Zr;
params.nZr = size(Zr, 2);
params.zmin = [];
params.zmax = [];
params.smin = [];%% Initializationdisp('Staring NSGA-III ...');empty_individual.Position = [];
empty_individual.Cost = [];
empty_individual.Rank = [];
empty_individual.DominationSet = [];
empty_individual.DominatedCount = [];
empty_individual.NormalizedCost = [];
empty_individual.AssociatedRef = [];
empty_individual.DistanceToAssociatedRef = [];pop = repmat(empty_individual, nPop, 1);
for i = 1:nPoppop(i).Position = unifrnd(VarMin, VarMax, VarSize);pop(i).Cost = CostFunction(pop(i).Position);
end% Sort Population and Perform Selection
[pop, F, params] = SortAndSelectPopulation(pop, params);%% NSGA-II Main Loopfor it = 1:MaxIt% Crossoverpopc = repmat(empty_individual, nCrossover/2, 2);for k = 1:nCrossover/2i1 = randi([1 nPop]);p1 = pop(i1);i2 = randi([1 nPop]);p2 = pop(i2);[popc(k, 1).Position, popc(k, 2).Position] = Crossover(p1.Position, p2.Position);popc(k, 1).Cost = CostFunction(popc(k, 1).Position);popc(k, 2).Cost = CostFunction(popc(k, 2).Position);endpopc = popc(:);% Mutationpopm = repmat(empty_individual, nMutation, 1);for k = 1:nMutationi = randi([1 nPop]);p = pop(i);popm(k).Position = Mutate(p.Position, mu, sigma);popm(k).Cost = CostFunction(popm(k).Position);end% Mergepop = [poppopcpopm]; %#ok% Sort Population and Perform Selection[pop, F, params] = SortAndSelectPopulation(pop, params);% Store F1F1 = pop(F{1});% Show Iteration Informationdisp(['Iteration ' num2str(it) ': Number of F1 Members = ' num2str(numel(F1))]);% Plot F1 Costsfigure(1);PlotCosts(F1);pause(0.01);end%% Resultsdisp(['Final Iteration: Number of F1 Members = ' num2str(numel(F1))]);
disp('Optimization Terminated.');
3、其他函数
3.1 AssociateToReferencePoint
function [pop, d, rho] = AssociateToReferencePoint(pop, params)Zr = params.Zr;nZr = params.nZr;rho = zeros(1, nZr);d = zeros(numel(pop), nZr);for i = 1:numel(pop)for j = 1:nZrw = Zr(:, j)/norm(Zr(:, j));z = pop(i).NormalizedCost;d(i, j) = norm(z - w'*z*w);end[dmin, jmin] = min(d(i, :));pop(i).AssociatedRef = jmin;pop(i).DistanceToAssociatedRef = dmin;rho(jmin) = rho(jmin) + 1;endend
3.2 Crossover
function [y1 y2] = Crossover(x1, x2)alpha = rand(size(x1));y1 = alpha.*x1+(1-alpha).*x2;y2 = alpha.*x2+(1-alpha).*x1;end
3.3 Dominate
function b = Dominates(x, y)if isstruct(x)x = x.Cost;endif isstruct(y)y = y.Cost;endb = all(x <= y) && any(x<y);end
3.4 GenerateReferencePoints
function Zr = GenerateReferencePoints(M, p)Zr = GetFixedRowSumIntegerMatrix(M, p)' / p;endfunction A = GetFixedRowSumIntegerMatrix(M, RowSum)if M < 1error('M cannot be less than 1.');endif floor(M) ~= Merror('M must be an integer.');endif M == 1A = RowSum;return;endA = [];for i = 0:RowSumB = GetFixedRowSumIntegerMatrix(M - 1, RowSum - i);A = [A; i*ones(size(B, 1), 1) B];endend
3.5 MOP2
function z = MOP2(x)n = numel(x);z1 = 1-exp(-sum((x-1/sqrt(n)).^2));z2 = 1-exp(-sum((x+1/sqrt(n)).^2));z = [z1 z2]';end
3.6 Mutate
function y = Mutate(x, mu, sigma)nVar = numel(x);nMu = ceil(mu*nVar);j = randsample(nVar, nMu);y = x;y(j) = x(j)+sigma*randn(size(j));end
3.7 NonDominatedSorting
function [pop, F] = NonDominatedSorting(pop)nPop = numel(pop);for i = 1:nPoppop(i).DominationSet = [];pop(i).DominatedCount = 0;endF{1} = [];for i = 1:nPopfor j = i+1:nPopp = pop(i);q = pop(j);if Dominates(p, q)p.DominationSet = [p.DominationSet j];q.DominatedCount = q.DominatedCount+1;endif Dominates(q.Cost, p.Cost)q.DominationSet = [q.DominationSet i];p.DominatedCount = p.DominatedCount+1;endpop(i) = p;pop(j) = q;endif pop(i).DominatedCount == 0F{1} = [F{1} i];pop(i).Rank = 1;endendk = 1;while trueQ = [];for i = F{k}p = pop(i);for j = p.DominationSetq = pop(j);q.DominatedCount = q.DominatedCount-1;if q.DominatedCount == 0Q = [Q j];q.Rank = k+1;endpop(j) = q;endendif isempty(Q)break;endF{k+1} = Q;k = k+1;endend
3.8 NormalizePopulation
function [pop, params] = NormalizePopulation(pop, params)params.zmin = UpdateIdealPoint(pop, params.zmin);fp = [pop.Cost] - repmat(params.zmin, 1, numel(pop));params = PerformScalarizing(fp, params);a = FindHyperplaneIntercepts(params.zmax);for i = 1:numel(pop)pop(i).NormalizedCost = fp(:, i)./a;endendfunction a = FindHyperplaneIntercepts(zmax)w = ones(1, size(zmax, 2))/zmax;a = (1./w)';end
3.9 PerformScalarizing
function params = PerformScalarizing(z, params)nObj = size(z, 1);nPop = size(z, 2);if ~isempty(params.smin)zmax = params.zmax;smin = params.smin;elsezmax = zeros(nObj, nObj);smin = inf(1, nObj);endfor j = 1:nObjw = GetScalarizingVector(nObj, j);s = zeros(1, nPop);for i = 1:nPops(i) = max(z(:, i)./w);end[sminj, ind] = min(s);if sminj < smin(j)zmax(:, j) = z(:, ind);smin(j) = sminj;endendparams.zmax = zmax;params.smin = smin;endfunction w = GetScalarizingVector(nObj, j)epsilon = 1e-10;w = epsilon*ones(nObj, 1);w(j) = 1;end
3.10 SortAndSelectPopulation
function [pop, F, params] = SortAndSelectPopulation(pop, params)[pop, params] = NormalizePopulation(pop, params);[pop, F] = NonDominatedSorting(pop);nPop = params.nPop;if numel(pop) == nPopreturn;end[pop, d, rho] = AssociateToReferencePoint(pop, params);newpop = [];for l = 1:numel(F)if numel(newpop) + numel(F{l}) > nPopLastFront = F{l};break;endnewpop = [newpop; pop(F{l})]; %#okendwhile true[~, j] = min(rho);AssocitedFromLastFront = [];for i = LastFrontif pop(i).AssociatedRef == jAssocitedFromLastFront = [AssocitedFromLastFront i]; %#okendendif isempty(AssocitedFromLastFront)rho(j) = inf;continue;endif rho(j) == 0ddj = d(AssocitedFromLastFront, j);[~, new_member_ind] = min(ddj);elsenew_member_ind = randi(numel(AssocitedFromLastFront));endMemberToAdd = AssocitedFromLastFront(new_member_ind);LastFront(LastFront == MemberToAdd) = [];newpop = [newpop; pop(MemberToAdd)]; %#okrho(j) = rho(j) + 1;if numel(newpop) >= nPopbreak;endend[pop, F] = NonDominatedSorting(newpop);end
3.11 UpdateIdealPoint
function zmin = UpdateIdealPoint(pop, prev_zmin)if ~exist('prev_zmin', 'var') || isempty(prev_zmin)prev_zmin = inf(size(pop(1).Cost));endzmin = prev_zmin;for i = 1:numel(pop)zmin = min(zmin, pop(i).Cost);endend